-
Archives
- June 2026
- November 2025
- October 2025
- September 2024
- October 2023
- August 2023
- January 2023
- December 2022
- October 2022
- September 2022
- June 2022
- March 2021
- August 2020
- June 2020
- December 2019
- October 2019
- September 2019
- July 2019
- June 2019
- May 2019
- April 2019
- January 2019
- November 2018
- September 2018
- August 2018
- March 2018
- September 2017
- July 2017
- June 2017
- April 2017
- November 2016
- August 2016
- March 2016
- November 2015
- September 2015
- August 2015
- July 2015
- March 2015
- February 2015
- October 2014
- September 2014
- July 2014
- June 2014
- May 2014
- January 2014
- September 2013
- June 2013
- April 2013
- March 2013
- February 2013
- January 2013
- December 2012
- June 2011
-
Meta
Save the date!
Stealing the message that was sent to the Boston flow email list:
We’re excited to share that NECyto’s 2026 Annual Meeting will be held on November 5th, 2026 at Moderna!
As we begin planning, we would greatly appreciate your input to shape this year’s meeting. Your feedback will help us create an agenda with topics and formats that are most valuable to our community.
Please take a few minutes to complete this short survey:
https://forms.gle/gRnfJ3e9mtjwbiUH9
Thank you in advance for your time and input. We look forward to seeing you in November!
Best,
The NECyto Committee
We’re excited to share that NECyto’s 2026 Annual Meeting will be held on November 5th, 2026 at Moderna!
As we begin planning, we would greatly appreciate your input to shape this year’s meeting. Your feedback will help us create an agenda with topics and formats that are most valuable to our community.
Please take a few minutes to complete this short survey:
https://forms.gle/gRnfJ3e9mtjwbiUH9
Thank you in advance for your time and input. We look forward to seeing you in November!
Best,
The NECyto Committee
Posted in Uncategorized
Leave a comment
2 days to go!
Only 2 more days until the return of NECyto’s fall meeting! I know you have all been anxiously awaiting the itinerary, you will find it pasted below.
Please note, our hosts have asked that we use the entrance on Fulkerson Street (the star on the map) just around the corner from the Binney Street entrance.
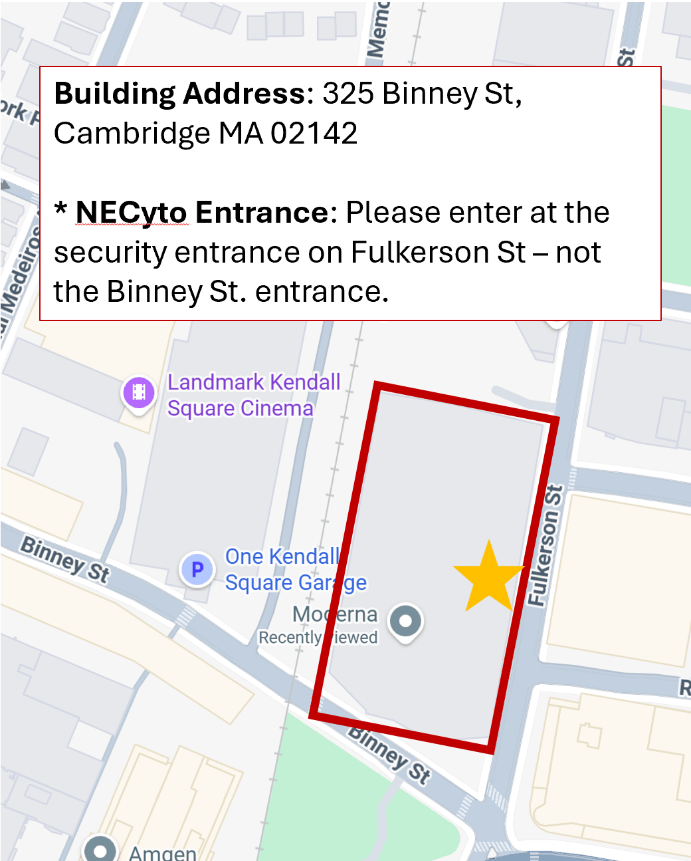
And our itinerary for the day, I look forward to seeing everyone!
–Mike

Posted in Uncategorized
Leave a comment
NECyto Speakers 2025

Hello flow community! I hope everyone had a fun Halloween, only 10 days until NECyto 2025 on Thursday, November 13! I’m pleased to announce our speaker lineup for this year, we will post the agenda and actual order of talks later this week:
Hsiao Hsuan Kuo, Moderna Tx: “Functional Profiling Using High Parameter Spectral Cytometry for Clinical Phase Studies in Vaccine Development”
Daniel Vocelle, St Jude Children’s Research Hospital: “Applications of Imaging Cytometry: From Technology to Discovery”
Spencer Shah, Broad Institute: “Exploring the use of imaging-based flow cytometers for functional genomics”
Mikhail Roshal, Memorial Sloan Kettering Cancer Center: “Myeloid neoplasms through the lens of high parameter flow cytometry”
Wayne Austin, Beam Tx: “Epitope engineering for in vivo enrichment of gene edited therapies”
To register, visit https://www.eventbrite.com/e/necyto-2025-tickets-1736133153999?aff=oddtdtcreator
We appreciate the hospitality of our hosts, Moderna. There are additional security registration steps required, so please register this week if you can!
All activities for the meeting will be at the Moderna facility at 325 Binney Street, Cambridge MA. Registration and breakfast starts at 8, and we will be heading back to MEX on Main Street, Cambridge, for the “Impromptu BBG” after the meeting.
On behalf of the organizing committee, we hope you can join us, and meet the new committee members!
Mike Waring
President
Posted in Uncategorized
Leave a comment
Welcome Back: NECyto 2025!
Another year has passed, and the annual Fall meeting is back! We are all settled into our new spaces, and this years meeting will also be in a new space: the Moderna Facility on Binney Street in Cambridge MA. Join us on November 13th for 5 talks and a vendor fair. We will be releasing the speaker list next week, as we finalize the talk titles.
I was cleaning out some papers and came across the agenda for our “Immunity and Immune Function in AIDS–Cytometric Methods” meeting from October 5, 2004, if you want to see what we were talking about 21 years ago click here!
Registration link is live! Go here to register!:
https://www.eventbrite.com/e/necyto-2025-tickets-1736133153999?aff=oddtdtcreator
We also have a few new committee members joining us this year, so we look forward to introducing them. There will be an “Impromptu BBG” after the meeting.
There will also be a “scheduled” BBG on Monday November 3, we hope you will join us for one or both of the upcoming events! Keep an eye on the mailing list for details (you can sign up on the “contact us” page).
Posted in Uncategorized
Leave a comment
Well THAT was a fast year!
Hello New England Cytometry users! I cant believe its been almost a year since the last post here, I had sent out notices on the mailing lists and social media that we would be skipping the fall meeting this year, but never posted it here! We are still getting settled into our new building for the Ragon Institute, and there are lots of changes. We do have an open position in the flow core, there is a technician level job posting here:
https://partners.taleo.net/careersection/ex/jobdetail.ftl?job=3301915
but we are also considering a more experienced position instead, so if you have 5+ years and would be interested in joining our team, let me know!
Monday October 7 we will have our monthly happy hour (or “BBG”) at MEX in Cambridge (500 Technology Square), across the street from the new building (600 Main Street). I’ll be happy to bring folks over in smaller groups to see the new space. Go to the “Contact us” page for the link to sign up for our distribution list if you arent on it already. Looking forward to seeing some of you soon!
Mike W
Posted in Uncategorized
Leave a comment
NECyto RECap
A HUGE Thank you to all of the attendees and vendors that joined us on October 12th for our fall meeting! Presentations are posted to the “Meetings” page, and it looked like everyone had a great time at MEX for the happy hour! The hostess said we were a really fun group and they’d be happy to host us again, so thank you everyone for being on your best behavior!
In over eleventy one months of running our social media accounts, I update half of them half as often as I should like, but I hope you like more than half the posts half as well as they deserve! We will try to post more in the coming year. We were thrilled to have Akos bring his camera, and he was busy all day! Enjoy the pictures on Facebook and Instagram, hopefully more to come. If you have any pictures to share, please send them our way or post them to the page!
Instagram (newenglandcytometry)
FaceBook (NECyto)
Twitter (@NECytometry)
LinkedIn (New England Cytometry)
ISAC
And don’t forget to sign up for the email list to hear about upcoming events!
https://elist.tufts.edu/sympa/info/hsfc to sign up or check the archives.

Posted in Uncategorized
Leave a comment
Two more days!
Our 2023 fall meeting is only 2 days away! We will be able to provide parking at the 800 Tech Square Garage for $10 for the day, look for the signs for NECyto.
Registration and check-in will be in the atrium of 100 Tech Square, vendors please do not enter before 7AM, attendees are invited to come for breakfast starting at 8AM.
Talks will begin at 9am in the first floor auditorium of 400 Tech Square across the street.
Attendees and vendors are invited to attend a post-meeting gathering at MEX in 500 Tech Square, lets keep the conversations going!
We are looking forward to seeing everyone!
Registration and check-in will be in the atrium of 100 Tech Square, vendors please do not enter before 7AM, attendees are invited to come for breakfast starting at 8AM.
Talks will begin at 9am in the first floor auditorium of 400 Tech Square across the street.
Attendees and vendors are invited to attend a post-meeting gathering at MEX in 500 Tech Square, lets keep the conversations going!
We are looking forward to seeing everyone!
Posted in Uncategorized
Leave a comment
Registration is open!
Registraion for the 2023 New England Cytometry User group meeting on Thursday, October 12, 2023 is now open! Use the button below to register on Eventbrite:
The meeting will be held at 400 Technology Square in Cambridge MA in the Ragon Institute’s Schwartz Auditorium for a day of talks, and a vendor fair in the atrium of 100/200 Tech Square. Breakfast will be available at 8. Speakers will include Sami Farhi (Broad Institute), Vera Tang (University of Ottawa), David Ng (University of Utah School of Medicine), Peter Lopez (NYU), Thomas Diefenbach (Ragon Institute), and a presentation from the Organizing Committee. Stick around afterwards for the “impromptu” happy hour. The full agenda will be posted to https://newenglandcytometry.com/meetings/ soon!
If you are retired and would like to attend, please contact us directly! Any vendors that are interested in additional sponsorship opportunities, also please reach out. Additional vendor information will be sent to registrants (venue access, shipping information).
Email us at mwaring@partners.org or new.england.cytometry@gmail.com.
We hope to see you there!
NECyto Organizing Committee
Posted in Uncategorized
Leave a comment
2023 New England Cytometry User group meeting!
Please save the date for the 2023 Fall NECyto meeting, to be held on Thursday October 12, 2023 returning to the Ragon Institute at 400 Tech Square in Cambridge! We will be opening up the registration site in the next couple of weeks, along with announcing the speaker list, but I wanted to assure our community that the meeting is happening and we are looking forward to welcoming everyone to Tech Square one last time, before the big move across the street! Be sure to pick up this year’s t-shirt, having done the visible spectrum:

this year we’re going UV!

We hope you’ll be visible at the meeting!
Mike Waring
President, NECyto User Group
Director, Ragon Institute Imaging Core Facility
Posted in Uncategorized
Leave a comment
Sorting Below Zero in Diva
When setting your sort gates in Diva, there are some limitations that users need to be aware of that are not obvious, especially when it comes to gating on populations close to or below the axis.
I first encountered this issue when samples were either overcompensated, or (as discussed in the previous post) when small fluorescent particles pushed the baseline restore artificially negative, so that the population of interest was at or below 0. There is a minimum “upper limit” for your sorting gate, and if you set the region boundary below this, no events will be sorted (your sort rate and collected events will be 0, even though you see events in the gate on the plots). This is true for ANY gate in the hierarchy, so if your “sort gate” is a descendant of a gate that is set too low, you will not register any sorted events.

Setting the gate any lower than shown for APC will result in no events being sorted.
The next limitation to highlight is that Diva cannot distinguish a LOWER boundary that is less than this cutoff—if you are sorting below this minimum limit, then ANYTHING below that is considered “in the gate” regardless of what lower boundary you set. In this example, region P5 is set around only the orange population, but in the post sort analysis on the right, you can see the green population was ALSO sorted.

Lastly, if you were going to try to sort just the green population, you could set at least one point of the gate above that “minimum threshold” value to get around that “minimum threshold” and get events to sort, and exclude the orange population from your gate, but you would end up still sorting both as it will sort anything below that minimum threshold value regardless of if it is gated or not. Again, post sort shows that even though only the green was gated, both green and orange populations are present in the post-sort sample.

We have found this to be true on both the Arias as well as the S6—The S6 allows for up to 16 gating levels vs. the Aria’s limit of 8, but still cannot distinguish gating below 0.
(I set this up with calibrite beads, FITC is green, PE is orange, then unstained and APC are the blue. I purposely overcompensated FITC from APC, and PE from APC, to push the populations below 0 but still be able to distinguish them.)
Posted in Uncategorized
Leave a comment
